ITS2 PCR 5/21/19
P2 P5 P6 P7 P8 P9 / P10 P11 P12 P13 P14 P15 P16
Procedure
PCR was performed on samples that have not been used before, these samples are from aliquoites of DNA extractions becuase the original coral samples are not available
Samples P2 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 were used in this PCR (polimerase chain reaction) reaction
First A Master Mix of water, 2x mytaq mix, ITS2 Primer Reverse (concentration: 10 micromolar), ITS2 Primer Forward (concentration: 10 micromolar)
| Component | Volume for 13 reactions(+5% error) |
|---|---|
| Water | 122.85 micoleters |
| 2x mytaq mix | 170.625 micoleters |
| ITS2 Primer Reverse (concentration: 10 micromolar) | 13.65 micoleters |
| ITS2 Primer Forward (concentration: 10 micromolar) | 13.65 micoleters |
24 micoleters of this master mix was added to 13 tubes For the 2 controls 12.5 micoleters water and 12.5 micoleters mytaq mix were added
1 microleter of the DNA samples was added to the coresponding tube and 1 microleter of water was added to two tubes for the controls
the samples were in two seperate rows of tubes
- Row 1: P2 P5 P6 Control P7 P8 P9 Control
- Row 2: P10 P11 P12 Control P13 P14 P15 P16
The samples were then placed in the thermocycler set to
| Step | Temperature | Duration | cycles |
|---|---|---|---|
| 1 | 92°C | 3:00 | 1 |
| 2 | 92°C | 0:30 | 35 |
| 3 | 52°C | 0:40 | 35 |
| 4 | 72°C | 0:30 | 35 |
| 5 | 72°C | 5:00 | 1 |
| 6 | 10°C | hold | - |
these amplified DNA samples were then tested by making a gel
to create the Gel the following steps were taken
1.5g agarose and 75ml 1X TAE buffer were mixed to make a 2% agarose 75ml gel mix the mix was microwaved and swirled until there were no flakes then 1 microliter Gelgreen was added
the mix was left to cool for 5 minutes then poured into the gel mold the gel was left to harden for 30 minutes
once hardened enough Used TAE buffer was poured into the gel box to cover the mold
5 microliter aliquots of DNA samples P2 P5 P6 P7 P8 P9 P10 P11 P12 P13 P14 P15 P16 and the three controls were made 1 microliter of 5x loading dye was added to each
First the HyperLadder 100bp was added to the first well of the mold then the DNA samples were each added to the wells in the following order
| Ladder | P2 | P5 | P6 | Control | P7 | P8 | P9 | Control |
| Ladder | P10 | P11 | P12 | Control | P13 | P14 | P15 | P16 |
The voltage was set to 100V and run for 60 minutes
These are the resulting Gels
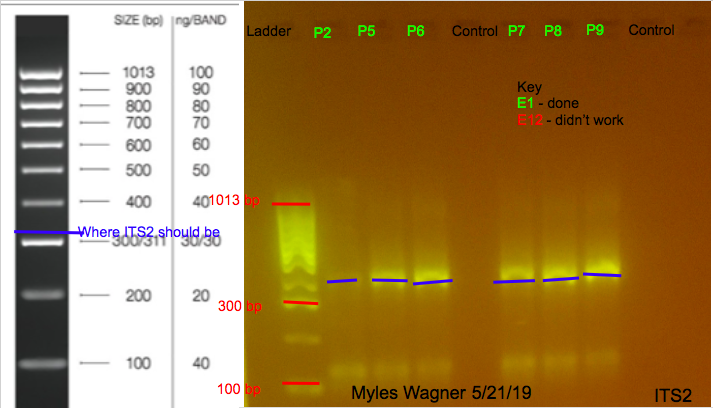

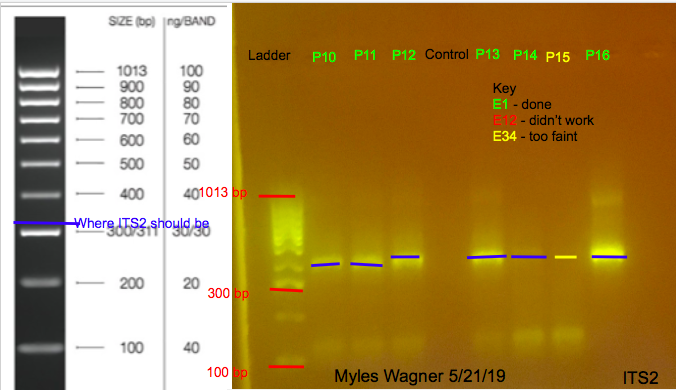
Notes
ALmost all worked Since ITS2 is found on the symbionts this may show that the Plexaura corals have more symbiont present that Eunicea