Learning to make Phylogentic trees #2
COI tree
Procedure
First a document was made with all the sequences for the Gene I was looking at
In this case the marker COI was used for all of the species of coral that I have been using in the PCRs
here are a list of all the codes used; COI sequences
These were then shortened to be the samelength and aligned using the program AliView https://ormbunkar.se/aliview/
Then using the website Cipres https://www.phylo.org the aligned sequences were analysed using the RAxML-HPC2 on XSEDE tool to create a Phylogenetic tree using maximum likelihood
This produced the following code
(Eunicea tourneforti(Eunicea tourneforti(((Eunicea succinea,Eunicea clavigera),Eunicea calyculata),(((((Eunicea fusca,Eunicea calyculata),Eunicea fusca),Eunicea fusca),(((Eunicea flexuosa,(Eunicea flexuosa,Eunicea flexuosa)),(Eunicea flexuosa,(Eunicea laxispica,Eunicea flexuosa))),Eunicea tourneforti)),Eunicea tourneforti))),Eunicea calyculata);
the names were then adjusted using the following codes
| Code | Species |
|---|---|
| MK153456.1 | Eunicea succinea |
| MK153476.1 | Eunicea flexuosa |
| MK153442.1 | Eunicea flexuosa |
| MK153329.1 | Eunicea flexuosa |
| MK153303.1 | Eunicea flexuosa |
| MK153374.1 | Eunicea flexuosa |
| MK153441.1 | Eunicea laxispica |
| MK153417.1 | Eunicea tourneforti |
| MK153414.1 | Eunicea tourneforti |
| MK153408.1 | Eunicea tourneforti |
| MK153376.1 | Eunicea tourneforti |
| MK153406.1 | Eunicea fusca |
| MK153362.1 | Eunicea fusca |
| MK153308.1 | Eunicea fusca |
| MK153384.1 | Eunicea clavigera |
| MK153378.1 | Eunicea calyculata |
| MK153334.1 | Eunicea calyculata |
| MK153432.1 | Eunicea calyculata |
finally using the site ITOL https://itol.embl.de a tree was made:
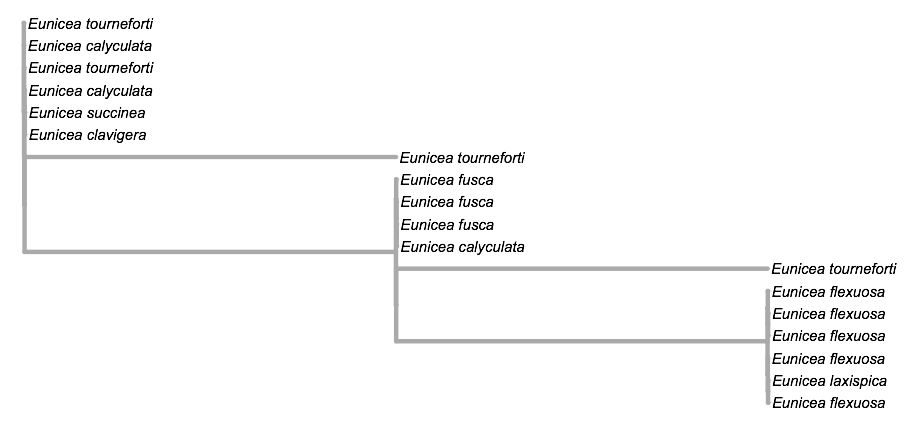
This tree can be shown in other ways as well
ignoring branch length:
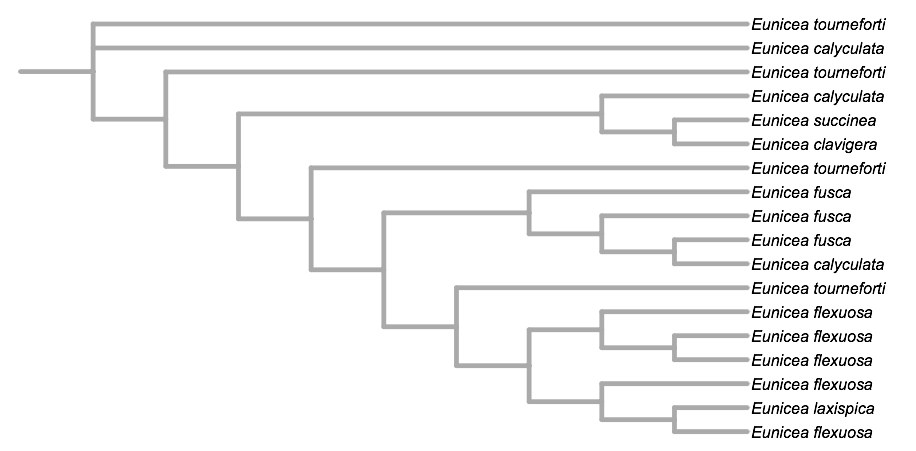
Circular:
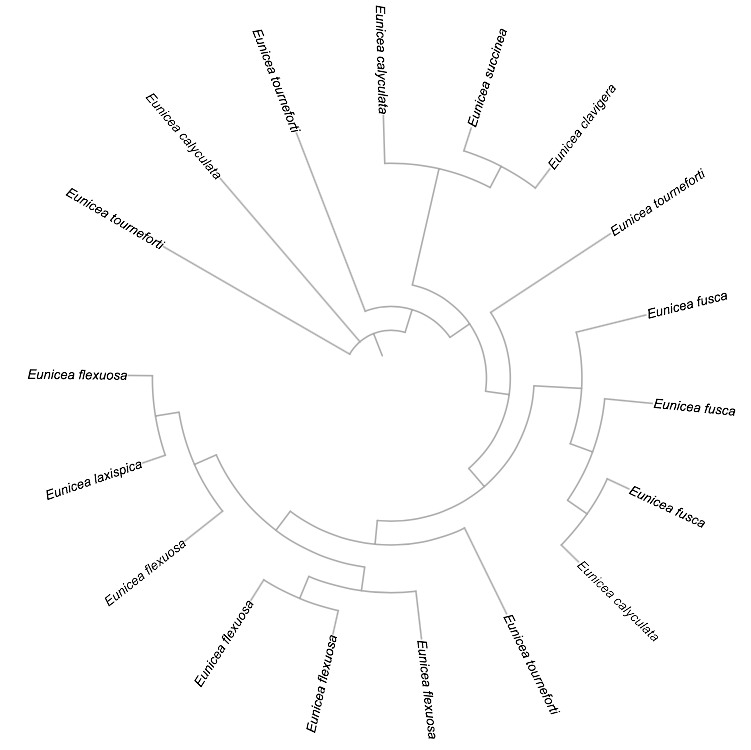
Uprooted:
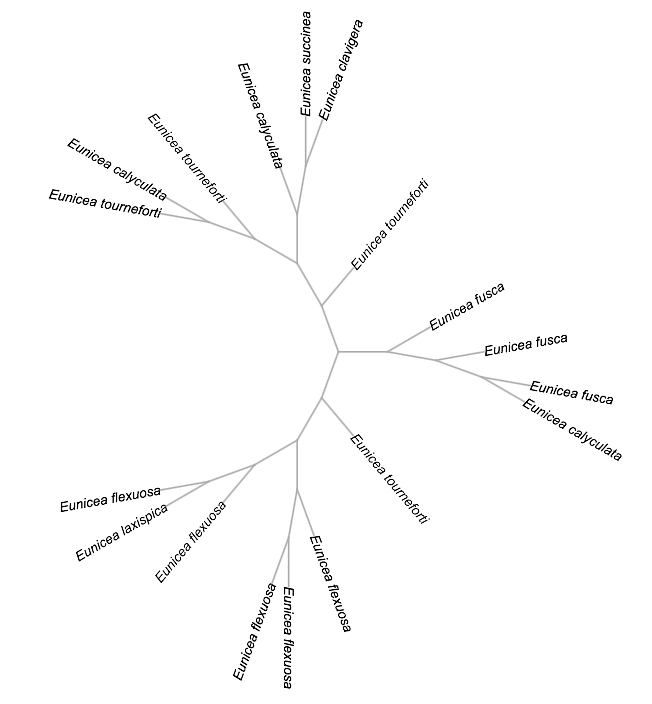
Analysis
This tree shows the maximum liklihood tree for the COI marker This tree has multiples of the same species to see how that effects the tree it is still only done with one marker
That is why in the study I am conducting many markers will be used and multiples of each species as well