CA6 PCR #4 8/5/19 - Temperature Gradient
E7
Procedure
Samples E7 were used in this PCR (polimerase chain reaction) reaction
First A Master Mix of water, 2x mytaq mix, CA6.38R Primer (concentration: 10 micromolar), CA6.38L Primer (concentration: 10 micromolar)
| Component | Volume for 14 reactions(+5% error) |
|---|---|
| Water | 66.15 micoleters |
| 2x mytaq mix | 91.9 micoleters |
| CA6.38R Primer (concentration: 10 micromolar) | 7.35 micoleters |
| CA6.38L Primer (concentration: 10 micromolar) | 7.35 micoleters |
| BAS (Borine Serum Albumin) | 1 microliter |
the BAS is at twice the concentration as last time
24 micoleters of this master mix was added to 14 tubes For the 2 negative controls 12.5 micoleters water and 12.5 micoleters mytaq mix were added there was no positive control was available
1 microleter of the DNA samples was added to the coresponding tube
This PCR was set to 30 cycles Previous PCRs were showing up smeared and unclear. A temperature gradient was set up to try to determine a temperature where CA6 works best
| Step | Temperature | Duration | cycles |
|---|---|---|---|
| 1 | 92°C | 3:00 | 1 |
| 2 | 92°C | 0:30 | 30 |
| 3 | Gradient | 1:00 | 30 |
| 4 | 72°C | 0:15 | 30 |
| 5 | 72°C | 2:00 | 1 |
| 6 | 4°C | hold | - |
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
| 54.7°C | 55.4°C | 56.2°C | 57.2°C | 58.2°C | 59.3°C | 60.2°C | 61.0°C |
these amplified DNA samples were then tested by making a gel
to create the Gel the following steps were taken
1.5g agarose and 75ml 1X TAE buffer were mixed to make a 2% agarose 75ml gel mix the mix was microwaved and swirled until there were no flakes then 1 microliters Gelgreen was added
the mix was left to cool for 5 minutes then poured into the gel mold the gel was left to harden for 30 minutes
once hardened enough Used TAE buffer was poured into the gel box to cover the mold
5 microliter aliquots of DNA samples E7 and the two controls were made 1 microliter of 5x loading dye was added to each
First 3 microliters of the HyperLadder 100bp was added to the first well of the mold then the DNA samples were each added to the wells in the following order
| Ladder | E7 | E7 | E7 | E7 | E7 | E7 | E7 | E7 |
The voltage was set to 100V and run for 60 minutes
This is the resulting Gel
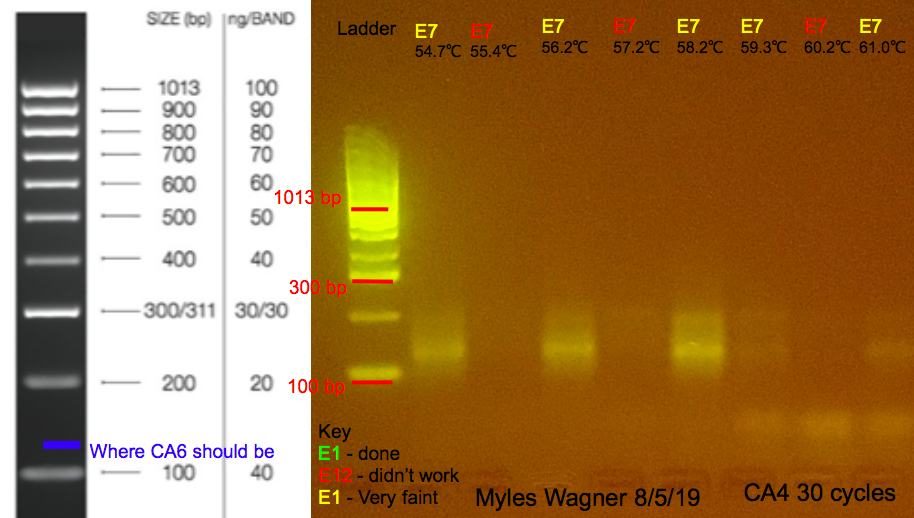
Notes
The PCR came out very different than expected. each well had the same mix in it just run at different temperatures through the Gradient. In the Gel it seemed to flip back and forth when really it should have been more graduel for which ones worked This PCR did not help much in determining the temperature that would work best for CA6
the temps that worked the best were 54.7°C, 56.2°C, and 58.2°C previously 56°C was used since that was the temperture suggested from previous liturature; (Santos, S.R., Guiterrez-Rodriguez, C., Lasker, H.R., Coffroth, M.A., 2003. Symbiodi- nium sp. associations in the gorgonian Pseudopterogorgia elisabethae in the Bahamas: high levels of genetic variability and population structure in symbi- otic dinoflagellates. Marine Biology 143, 111–120.)